  
Other potentially pathogenic fastidious GNR such as Capnocytophaga spp. 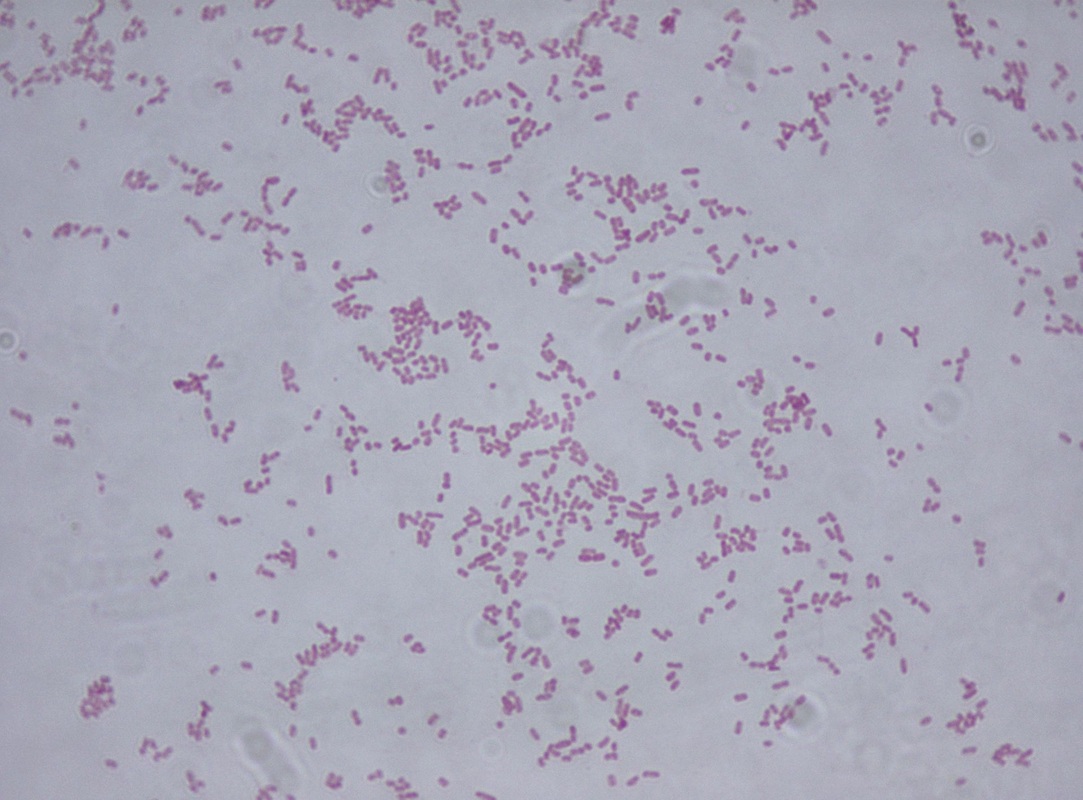
corrodens, and Kingella spp.), however, only a small set of isolates and species were investigated. The application of newer identification methods like matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) shows promising results regarding the identification of HACEK group members ( Haemophilus parainfluenzae, Aggregatibacter spp., Cardiobacterium spp., E. Most studies relied only on a subset of taxa of fastidious GNR or did not include clinical isolates under routine conditions. Commercially available identification systems such as VITEK 2 NH (bioMérieux, Marcy L’Etoile, France) only partially allow for accurate identification of this group of microorganisms, e.g., Eikenella corrodens, Kingella kingae and Cardiobacterium hominis. Identification of fastidious GNR by conventional methods is difficult and time-consuming because phenotypic characteristics such as growth factor requirements, fermentation and assimilation of carbohydrates, morphology, and staining behaviour are subject to variation and dependent on individual interpretation and expertise. Accurate identification of fastidious GNR is of concern when isolated from normally sterile body sites regarding guidance of appropriate antimicrobial therapy and patient management. Most of them are colonizers of the human oral cavity but they have been demonstrated to cause severe systemic infections like endocarditis, septicemia and abscesses, particularly in immunocompromised patients. They are isolated infrequently and consist of different taxa including Actinobacillus, Capnocytophaga, Cardiobacterium, Eikenella, Kingella, Moraxella, Neisseria, and Pasteurella. Fastidious GNR are slow-growing organisms, which generally require supplemented media or CO 2 enriched atmosphere and fail to grow on enteric media such as MacConkey agar. We conclude that 16S rRNA gene sequencing is an effective means for identification of fastidious GNR, which are not readily identified by conventional phenotypic methods.Īccurate identification of fastidious Gram-negative rods (GNR) is a challenge for clinical microbiology laboratories. We herein propose an efficient strategy for accurate identification of fastidious GNR in the clinical microbiology laboratory by integrating both conventional phenotypic methods and 16S rRNA gene sequence analysis. 73/158 (47%) of the isolates were not identified or misidentified. Compared to 16S rRNA gene sequencing as reference method, phenotypic identification correctly identified 64/158 (40%) isolates to species level, mainly Aggregatibacter aphrophilus, Cardiobacterium hominis, Eikenella corrodens, Pasteurella multocida, and 21/158 (13%) isolates correctly to genus level, notably Capnocytophaga sp. 16S rRNA gene homology analysis identified 148/158 (94%) of the isolates to species level, 9/158 (5%) to genus and 1/158 (1%) to family level. ResultsĪ total of 158 clinical isolates covering 20 genera and 50 species isolated from 1993 to 2010 were analyzed by comparing biochemical and 16S rRNA gene sequence analysis based identification. The aim of this study was to evaluate the use of molecular methods, e.g., 16S rRNA gene sequence analysis for identification of fastidious GNR in the clinical microbiology laboratory. Accurate identification of fastidious Gram-negative rods (GNR) by conventional phenotypic characteristics is a challenge for diagnostic microbiology. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
